Workflow Example 2 - Multiple Sites
Peter Levy
2026-03-03
workflow_multisite.RmdThis vignette illustrates the typical workflow for quality control of
met data from multiple sites simultaneously with metamet.
We assume that the basic steps have been carried out already: reading
the raw observation data, creating systematic metadata for the variables
and each site (described in Workflow
Example 1). This produces a single metamet object for
each site.
First, we load the metamet library, and read the data
for each of three sites from file.
here::i_am("vignettes/workflow_example.Rmd")
library(here)
library(metamet)
mm_amo <- readRDS(
file = here::here("data-raw/UK-AMO/UK-AMO_BM_mm_2023.rds")
)
mm_ebu <- readRDS(
file = here::here("data-raw/UK-EBU/UK-EBU_BM_mm_2023.rds")
)
mm_whm <- readRDS(
file = here::here("data-raw/UK-WHM/UK-WHM_BM_mm_2023.rds")
)This gives us all the half-hourly data for 2023 from each site. If we
want to process a shorter time period, we can subset as required with
subset_by_date. Here we look at two days just for
speed.
mm_amo <- subset_by_date(mm_amo, "2023-09-01", "2023-09-03")
mm_ebu <- subset_by_date(mm_ebu, "2023-09-01", "2023-09-03")
mm_whm <- subset_by_date(mm_whm, "2023-09-01", "2023-09-03")In order to combine data from different sites in a meningful way,
they all need to use the same naming convention. Here we convert two of
the sites to the ICOS convention using the
change_naming_convention function below (UK-AMO data are
already produced in this form).
mm_ebu <- change_naming_convention(mm_ebu, name_convention = "name_icos")
mm_whm <- change_naming_convention(mm_whm, name_convention = "name_icos")Although the base names of variable are now standardised across
sites, the specific variables measured will vary across sites. For
example, one site may have two replicate measurements of air temperature
TA_4_1_1 and TA_5_1_1, while another has a single replicate TA_1_1_1.
These cannot be combined as a single variable (or related variables) in
wide format. Also, wide format would require columns for the full set of
variables found across all sites, many of which will be empty for all
sites except one. The efficient solution is to convert to long format,
so all values are in a single column, with additional columns to
identify the site, timestamp, varaiable type and specific varaiable
name. We do this with the reshape_wide_to_long function,
shown below.
mm_amo <- reshape_wide_to_long(mm_amo)
mm_ebu <- reshape_wide_to_long(mm_ebu)
mm_whm <- reshape_wide_to_long(mm_whm)We can now see the structure of UK-AMO data
mm_amo$dt
#> Key: <site, TIMESTAMP, var_name>
#> site TIMESTAMP var_name value type name_icos
#> <char> <POSc> <fctr> <num> <char> <char>
#> 1: UK-AMO 2023-09-01 TS 11.9883733 temperature TS
#> 2: UK-AMO 2023-09-01 SWC 75.7355696 soil moisture SWC
#> 3: UK-AMO 2023-09-01 G 1.2816722 energy flux G
#> 4: UK-AMO 2023-09-01 WTD -0.3445917 height WTD
#> 5: UK-AMO 2023-09-01 WTD_4_1_1 NA height WTD
#> ---
#> 3585: UK-AMO 2023-09-03 NDVI_649OUT_5_1_1 NA energy flux NDVI_649OUT
#> 3586: UK-AMO 2023-09-03 NDVI_797IN_5_1_1 NA energy flux NDVI_797IN
#> 3587: UK-AMO 2023-09-03 NDVI_797OUT_5_1_1 NA energy flux NDVI_797OUT
#> 3588: UK-AMO 2023-09-03 LWS_4_1_1 360.6000000 arbitrary LWS
#> 3589: UK-AMO 2023-09-03 LWS_4_1_2 285.0000000 arbitrary LWS
#> qc validator ref
#> <num> <char> <num>
#> 1: 0 auto 1.173913e+01
#> 2: 0 auto 3.360330e+01
#> 3: 0 auto -6.467518e-14
#> 4: 0 auto 3.360330e+01
#> 5: 1 auto 3.360330e+01
#> ---
#> 3585: 1 auto 0.000000e+00
#> 3586: 1 auto 0.000000e+00
#> 3587: 1 auto 0.000000e+00
#> 3588: 0 auto 8.707268e+01
#> 3589: 0 auto 8.707268e+01and see it is identical to the structure of UK-EBU (and UK-WHM) data.
mm_ebu$dt
#> Key: <site, TIMESTAMP, var_name>
#> site TIMESTAMP var_name value type name_icos qc
#> <char> <POSc> <fctr> <num> <char> <char> <num>
#> 1: UK-EBU 2023-09-01 RH_1_1_1 90.100 humidity RH 0
#> 2: UK-EBU 2023-09-01 TA_1_1_1 9.320 temperature TA 0
#> 3: UK-EBU 2023-09-01 LWS_7_1_1 286.500 arbitrary LWS 0
#> 4: UK-EBU 2023-09-01 PA_1_1_1 98.700 pressure PA 0
#> 5: UK-EBU 2023-09-01 P_1_1_1 0.000 precipitation P 0
#> ---
#> 1839: UK-EBU 2023-09-03 TS_5_3_1 14.930 temperature TS 0
#> 1840: UK-EBU 2023-09-03 TS_5_4_1 14.360 temperature TS 0
#> 1841: UK-EBU 2023-09-03 G_5_1_1 0.171 energy flux G 0
#> 1842: UK-EBU 2023-09-03 G_5_2_1 -3.032 energy flux G 0
#> 1843: UK-EBU 2023-09-03 WTD_5_1_1 -0.576 height WTD 0
#> validator ref
#> <char> <num>
#> 1: auto 88.06677
#> 2: auto 11.23692
#> 3: auto 88.06677
#> 4: auto 97.53844
#> 5: auto 0.00000
#> ---
#> 1839: auto 13.84093
#> 1840: auto 13.84093
#> 1841: auto 0.00000
#> 1842: auto 0.00000
#> 1843: auto 32.56382Given this structure, we can simply append all the rows for all data
tables to create a single metamet object.
mm <- rbind_metamet(
mm_amo,
l_dt = list(mm_amo$dt, mm_ebu$dt, mm_whm$dt),
l_dt_meta = list(mm_amo$dt_meta, mm_ebu$dt_meta, mm_whm$dt_meta),
l_dt_site = list(mm_amo$dt_site, mm_ebu$dt_site, mm_whm$dt_site)
)We can now plot all the variables of a given type for all the types, identifying site by colour or separate panels (facets). Below we plot soil temperature against time; the solid black line shows the reference data, which in this case is ERA5 reanalysis data for the grid cell containing the site.
p <- ggplot(
mm$dt[name_icos == "TS", ],
aes(TIMESTAMP, value, colour = var_name)
)
p <- p + geom_line(aes(y = ref), colour = "black")
p <- p + geom_point()
p <- p + facet_wrap(~site)
print(p)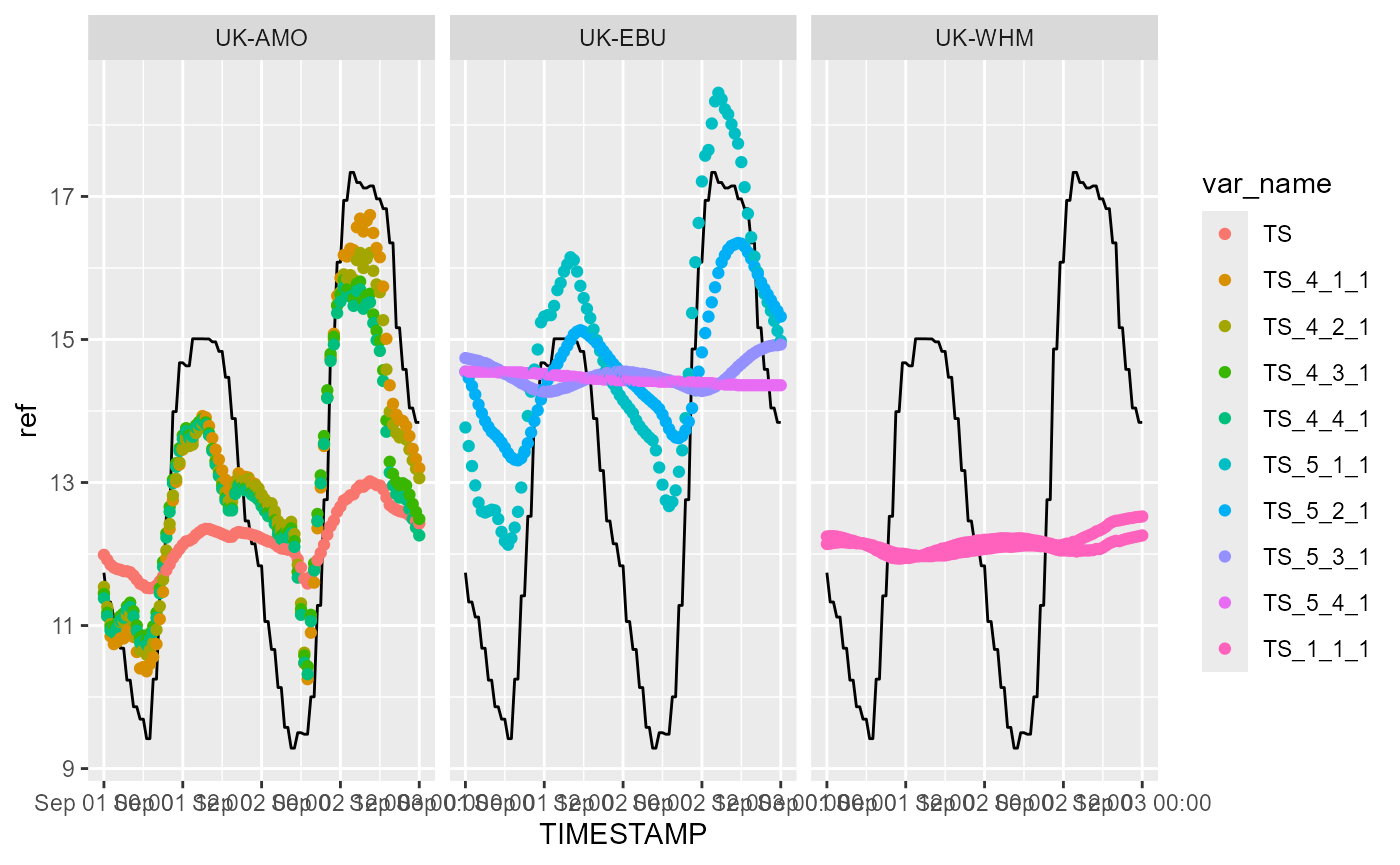
Below we plot PPFD (photosynthetic photon flux density, commonly ‘PAR’) against time; the solid black line shows the ERA5 reference data.
p <- ggplot(
mm$dt[name_icos == "PPFD_IN", ],
aes(TIMESTAMP, value, colour = var_name)
)
p <- p + geom_line(aes(y = ref), colour = "black")
p <- p + geom_line(alpha = 1.0)
p <- p + facet_wrap(~site)
print(p)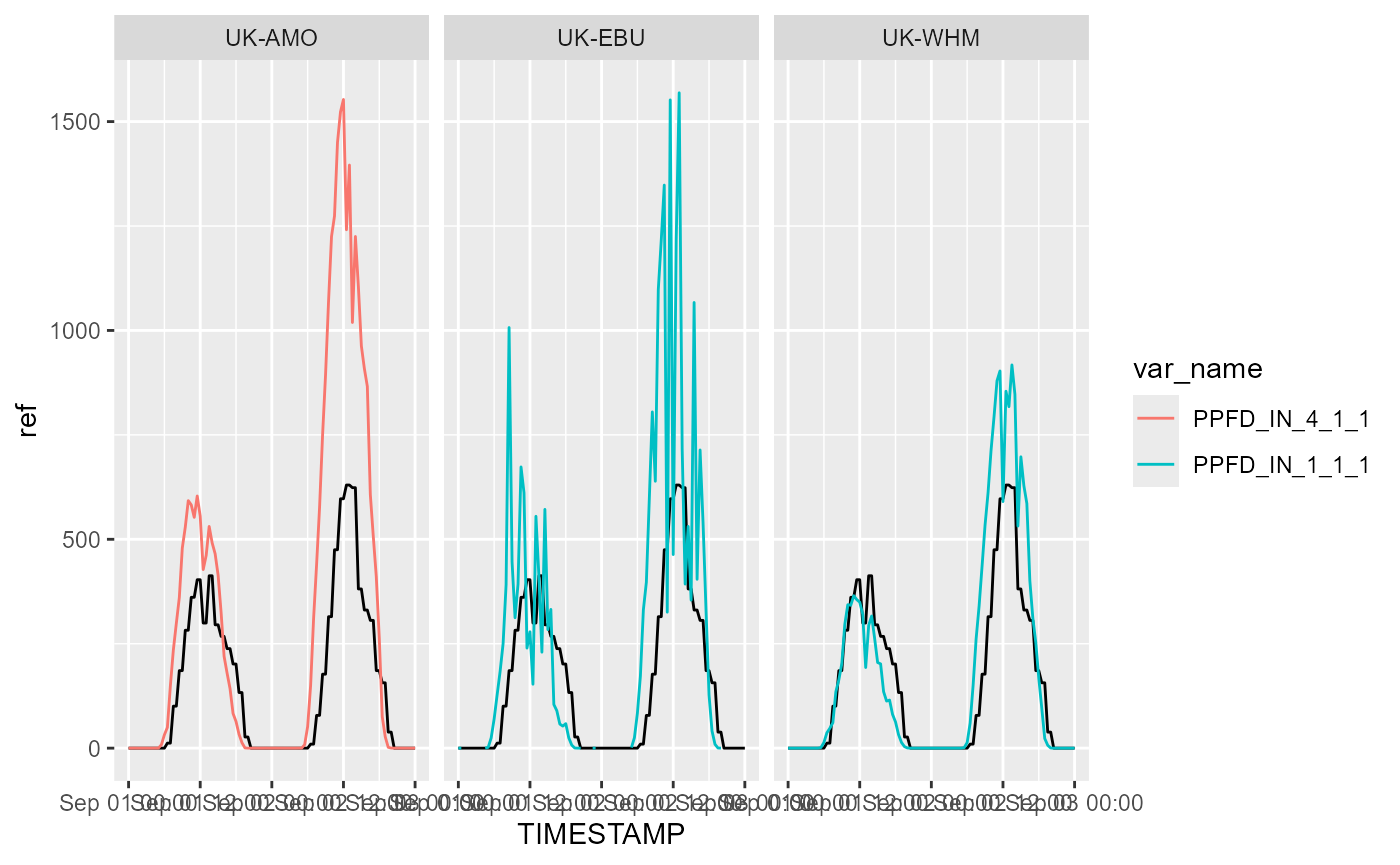
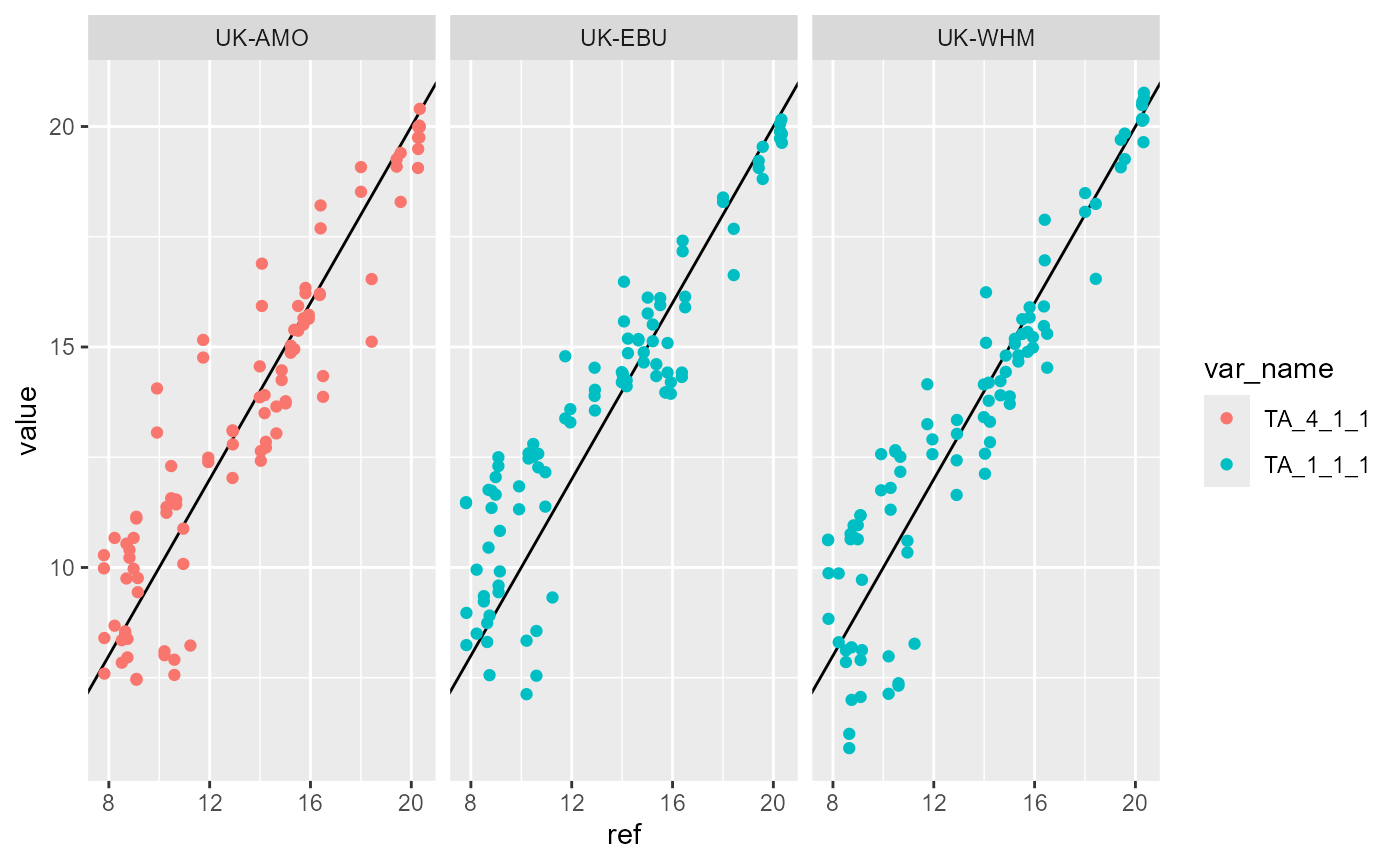
We can also plot all variable against the reference (ERA5) data to check for anomalous deviations from the expected relationship.
p <- ggplot(
mm$dt[name_icos == "TA", ],
aes(ref, value, colour = var_name)
)
p <- p + geom_abline()
p <- p + geom_point()
p <- p + facet_wrap(~site)
print(p)